DAG¶
Copying¶
DAG.copy() |
Return a copy of the current DAG. |
DAG.rename_nodes(name_map, VT]) |
Rename the nodes in this graph according to name_map. |
DAG.induced_subgraph(nodes) |
Return the induced subgraph over only nodes |
Information about nodes¶
DAG.parents_of(nodes, Set[Hashable]]) |
Return all nodes that are parents of the node or set of nodes nodes. |
DAG.children_of(nodes, Set[Hashable]]) |
Return all nodes that are children of the node or set of nodes nodes. |
DAG.neighbors_of(nodes, Set[Hashable]]) |
Return all nodes that are adjacent to the node or set of nodes node. |
DAG.markov_blanket_of(node) |
Return the Markov blanket of node, i.e., the parents of the node, its children, and the parents of its children. |
DAG.ancestors_of(nodes) |
Return the ancestors of nodes. |
DAG.descendants_of(nodes, Set[Hashable]]) |
Return the descendants of node. |
DAG.indegree_of(node) |
Return the indegree of node. |
DAG.outdegree_of(node) |
Return the outdegree of node. |
DAG.incoming_arcs(node) |
Return all arcs with target node. |
DAG.outgoing_arcs(node) |
Return all arcs with source node. |
DAG.incident_arcs(node) |
Return all arcs with node as either source or target. |
Graph modification¶
DAG.add_node(node) |
Add node to the DAG. |
DAG.add_nodes_from(nodes) |
Add nodes to the graph from the collection nodes. |
DAG.remove_node(node[, ignore_error]) |
Remove the node node from the graph. |
DAG.add_arc(i, j[, check_acyclic]) |
Add the arc i -> j to the DAG |
DAG.add_arcs_from(arcs[, check_acyclic]) |
Add arcs to the graph from the collection arcs. |
DAG.remove_arc(i, j[, ignore_error]) |
Remove the arc i -> j. |
DAG.reverse_arc(i, j[, ignore_error, …]) |
Reverse the arc i -> j to i <- j. |
Graph properties¶
DAG.has_arc(source, target) |
Check if this DAG has an arc source -> target. |
DAG.sources() |
Get all nodes in the graph that have no parents. |
DAG.sinks() |
Get all nodes in the graph that have no children. |
DAG.reversible_arcs() |
Get all reversible (aka covered) arcs in the DAG. |
DAG.is_reversible(i, j) |
Check if the arc i -> j is reversible (aka covered), i.e., if 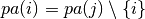 |
DAG.arcs_in_vstructures() |
Get all arcs in the graph that participate in a v-structure. |
DAG.vstructures() |
Get all v-structures in the graph, i.e., triples of the form (i, k, j) such that i->k<-j and i is not adjacent to j. |
DAG.triples() |
Return all triples of the form (i, j, k) such that i and k are both adjacent to j. |
DAG.upstream_most(s) |
Return the set of nodes which in s which have no ancestors in s. |
Ordering¶
DAG.topological_sort() |
Return a topological sort of the nodes in the graph. |
DAG.is_topological(order) |
Check that order is a topological order consistent with this DAG, i.e., if i->``j`` in the DAG, then i comes before j in the order. |
DAG.permutation_score(order) |
Return the number of “errors” in order with respect to the DAG, i.e., the number of times that i->``j`` in the DAG but i comes after j in order. |
Comparison to other DAGs¶
DAG.shd(other) |
Compute the structural Hamming distance between this DAG and the DAG other. |
DAG.shd_skeleton(other) |
Compute the structure Hamming distance between the skeleton of this DAG and the skeleton of the graph other. |
DAG.markov_equivalent(other[, interventions]) |
Check if this DAG is (interventionally) Markov equivalent to the DAG other. |
DAG.is_imap(other) |
Check if this DAG is an IMAP of the DAG other, i.e., all d-separation statements in this graph are also d-separation statements in other. |
DAG.is_minimal_imap(other[, certify, check_imap]) |
Check if this DAG is a minimal IMAP of other, i.e., it is an IMAP and no proper subgraph of this DAG is an IMAP of other. |
DAG.chickering_distance(other) |
Return the total number of edge reversals plus twice the number of edge additions/deletions required to turn this DAG into the DAG other. |
DAG.confusion_matrix(other[, rates_only]) |
Return the “confusion matrix” associated with estimating the CPDAG of other instead of the CPDAG of this DAG. |
DAG.confusion_matrix_skeleton(other) |
Return the “confusion matrix” associated with estimating the skeleton of other instead of the skeleton of this DAG. |
Separation statements¶
DAG.dsep(A, Hashable], B, Hashable], C, …) |
Check if A and B are d-separated given C, using the Bayes ball algorithm. |
DAG.dsep_from_given(A, C, …) |
Find all nodes d-separated from A given C. |
DAG.is_invariant(A, intervened_nodes[, …]) |
Check if the distribution of A given cond_set is invariant to an intervention on intervened_nodes. |
DAG.local_markov_statements() |
Return the local Markov statements of this DAG, i.e., those of the form i independent nondescendants(i) given the parents of i. |
Conversion to other formats¶
DAG.from_amat(amat) |
Return a DAG with arcs given by amat, i.e. |
DAG.to_amat([node_list]) |
Return an adjacency matrix for this DAG. |
DAG.from_nx(nx_graph) |
Convert a networkx DiGraph into a DAG. |
DAG.to_nx() |
Convert DAG to a networkx DiGraph. |
DAG.from_dataframe(df) |
Create a DAG from a dataframe, where the indices and columns are node names and a nonzero entry indicates the presence of an edge. |
DAG.to_dataframe([node_list]) |
Turn this DAG into a dataframe, where the indices and columns are node names and a nonzero entry indicates the presence of an edge. |
Conversion to other graphs¶
DAG.moral_graph() |
Return the (undirected) moral graph of this DAG, i.e., the graph with the parents of all nodes made adjacent. |
DAG.marginal_mag(latent_nodes[, relabel, new]) |
Return the maximal ancestral graph (MAG) that results from marginalizing out latent_nodes. |
DAG.cpdag() |
Return the completed partially directed acyclic graph (CPDAG, aka essential graph) that represents the Markov equivalence class of this DAG. |
DAG.interventional_cpdag(interventions[, cpdag]) |
Return the interventional essential graph (aka CPDAG) associated with this DAG. |
Chickering Sequences¶
DAG.resolved_sinks(other) |
Return the nodes in this graph which are “resolved sinks” with respect to the graph other. |
DAG.chickering_sequence(imap[, verbose]) |
Return a Chickering sequence from this DAG to an I-MAP imap. |
DAG.apply_edge_operation(imap[, seed_sink, …]) |
Identify an edge operation (covered edge reversal or edge addition) which decreases the Chickering distance from this DAG to imap. |
Directed Clique Trees¶
DAG.directed_clique_tree([verbose]) |
Return the directed clique tree associated with this DAG. |
DAG.contracted_directed_clique_tree() |
Return the contracted directed clique tree associated with this DAG. |
DAG.residuals() |
Return the residuals associated with this DAG. |
DAG.residual_essential_graph() |
Return the residual essential graph associated with this DAG. |
Intervention Design¶
DAG.optimal_fully_orienting_single_node_interventions([…]) |
Find the smallest set of interventions which fully orients the CPDAG into this DAG. |
DAG.greedy_optimal_single_node_intervention([…]) |
Greedily pick num_interventions single node interventions based on how many edges they orient. |
DAG.greedy_optimal_fully_orienting_interventions([cpdag]) |
Find a set of interventions which fully orients a CPDAG into this DAG, using greedy selection of the interventions. |